  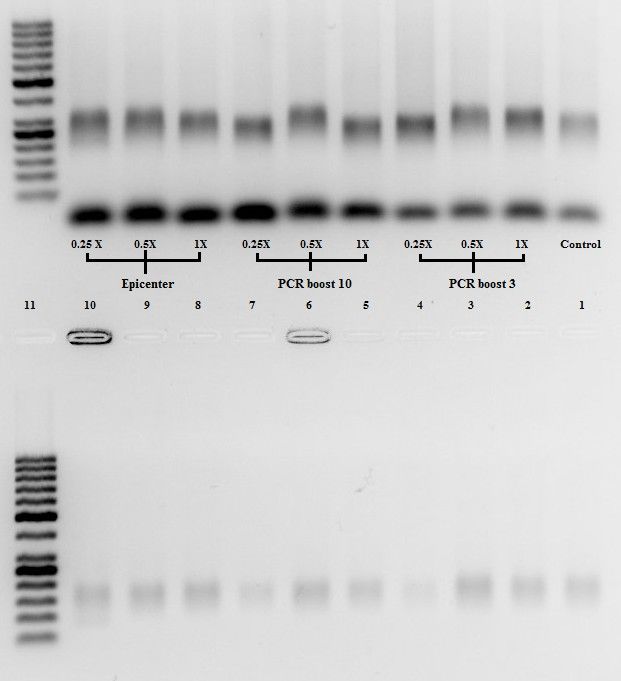
Incompletely elongated DNA strands are presumably the consequence of the pausing of DNA polymerase on the template, or of its premature termination. Chimeric DNA molecules seem to result from the annealing of incompletely extended primers to a heterologous target sequence during PCR. Such PCR products complicate and can mislead the MPRA data analysis as well as decrease the productivity of the approach, as the association of the same BC with different ROIs leads to the elimination of all such BCs and ROIs from the analysis. However, it has been previously shown that routine conventional PCR co-amplification of DNA sequences containing two variable motifs (in the case of MPRAs, BC and ROI) separated by a constant region frequently leads to formation of undesired chimeric molecules, from 5.4 to 30%. To identify all unique BC–ROI combinations, PCR amplification followed by the next-generation sequencing (NGS) is typically used. Importantly, the sequences of BCs and/or the associated DNA regions of interest (hereafter ROIs) in MPRA plasmid libraries are frequently not known a priori, as random oligonucleotides are used to clone these elements. Thus, the influence of each DNA sequence on the reporter expression can be measured by counting of the associated BC in the reporter RNA-seq data. To trace each individual DNA sequence under study, each molecule in the library is uniquely marked by a short (from a few to up to several dozen bp in length) barcode (BC), which is present within an untranslated region of the reporter gene. 
Briefly, a set of DNA sequences to assay are cloned in a plasmid vector outside of a reporter gene coding sequence, and the resulting MPRA library, which typically contains a high number of plasmid variants, is used to transfect the cells of interest. enhancers and promoters) or their mutant variants. Massively parallel reporter assays (MPRAs) allow high-throughput functional analysis of different DNA regulatory elements (e.g. In addition, we found that there is a negligible difference in performance of emulsion and conventional PCR approaches performed with the identified settings. We have identified PCR parameters that ensure synthesis of specific (non-chimeric) products from highly diverse MPRA plasmid libraries. Using specific MPRA libraries as templates, we found that the two-round amplification of the BC–ROI regions with a very low initial template amount, an elongated extension step, and a specific number of PCR cycles result in as low as 0.30 and 0.32% of chimeric products for emulsion and conventional PCR approaches, respectively. To identify settings that minimize formation of chimeric products we tested a number of PCR amplification parameters, such as conventional and emulsion types of PCR, one- or two-round amplification strategies, amount of DNA template, number of PCR cycles, and the duration of the extension step. However, chimeric DNA molecules formed on templates with identical spacer fragment during the amplification process may substantially hamper the identification of genuine BC–ROI combinations, and as a result lower the performance of the assays. Typically, this is done by PCR amplification of the BC–ROI regions with flanking primers, followed by next-generation sequencing (NGS) of the products. In the latter case, it is necessary to identify these combinations before performing functional experiments. The sequences of BC–ROI combinations present in the libraries may be either known a priori or not. The assays are based on construction of highly diverse plasmid libraries containing two variable fragments, a region of interest (a sequence under study ROI) and a barcode (BC) used to uniquely tag each ROI, which are separated by a constant spacer sequence. Massively parallel reporter assays (MPRAs) enable high-throughput functional evaluation of various DNA regulatory elements and their mutant variants. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
